At the probe station you capture one IV curve per device, one device per die, one die per wafer. The natural place to store the width and length is in the filename, but the moment a second operator runs the same experiment the abbreviations diverge (R_500_10.csv vs res_w500nm_l10um.csv), and collecting all the width variants for a single die later requires either grep gymnastics or a separate mapping spreadsheet. Tagging each file at upload time solves this: the physical dimensions travel with the data, not in its name.
In the previous notebook we generated a GDS layout with three resistance test structures of different widths and uploaded it to DataLab.
In a real experiment you would now probe each structure on a wafer using a four-probe measurement to get IV curves, then extract resistance from the slope. Here we simulate that process: for each combination of wafer, die, and device, we compute a synthetic IV curve using Ohm's law, add realistic noise and defects, and upload the result with provenance tags.
The tags on each file are what make the data queryable later. Each file carries: the project name, your username, the wafer ID, the die coordinates, the cell name (which encodes the width), the device instance name, the physical length, and the physical width. The analysis notebooks will use these tags to group measurements and extract sheet resistance.
Setup¶
import getpass
from contextlib import suppress
from functools import cache
from itertools import product
import gdsfactory as gf
import gfhub
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from gdsfactory.gpdk import PDK
from tqdm.notebook import tqdm
PDK.activate()
np.random.seed(42)
client = gfhub.Client()
user = getpass.getuser()
print(f"Running as user: {user}")
Running as user: runner
The measurement model¶
Each IV measurement sweeps current from 0 to 10 mA and records voltage. The resistance of each structure follows R = Rs × L / W, where Rs is the sheet resistance (100 Ω/sq for this simulation), L is the structure length, and W is the width. Voltage is then just V = R × I (Ohm's law).
To make the data realistic, we add three layers of variation:
- Wafer-to-wafer variation: up to ±20% multiplicative shift, constant across all dies on one wafer. This mimics run-to-run process variation.
- Device-to-device variation: up to ±5% additional random factor per device, representing local within-die non-uniformity.
- Measurement noise: 5% random multiplicative noise on each voltage sample.
- Defects: a small fraction of devices are simulated as open circuits (R = 1 GΩ) or short circuits (R = 0), which the analysis pipeline will need to handle.
def iv_resistance(resistance_ohm: float, current_mA: np.ndarray) -> np.ndarray:
"""Return voltage in mV for a given resistance and current sweep (Ohm's law)."""
return resistance_ohm * current_mA # mV = Ω × mA
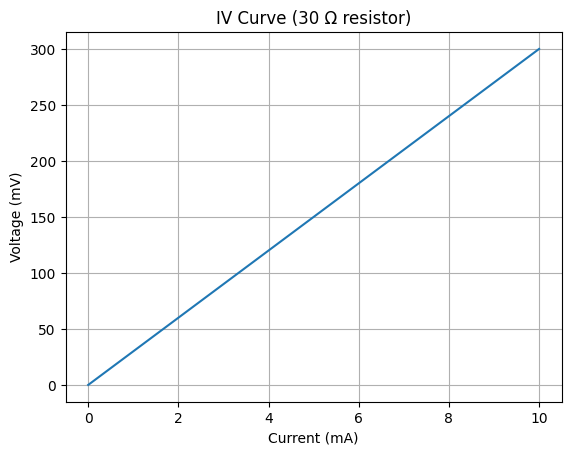
# Show a sample IV curve for a 30 Ω resistor
df_demo = pd.DataFrame(
{
"current_mA": (i := np.linspace(0, 10, 21)),
"voltage_mV": iv_resistance(resistance_ohm=30, current_mA=i),
}
)
plt.plot(df_demo.current_mA, df_demo.voltage_mV)
plt.title("IV Curve (30 Ω resistor)")
plt.xlabel("Current (mA)")
plt.ylabel("Voltage (mV)")
plt.grid(True)
plt.show()
Wafer and die definitions¶
We simulate three wafers and a 5×5 grid of dies with the four corner dies removed. The die coordinates range from (-2,-2) to (2,2).
wafer_ids = ["wafer1", "wafer2"]
dies = [
{"x": x, "y": y}
for y in range(-1, 1)
for x in range(-1, 1)
]
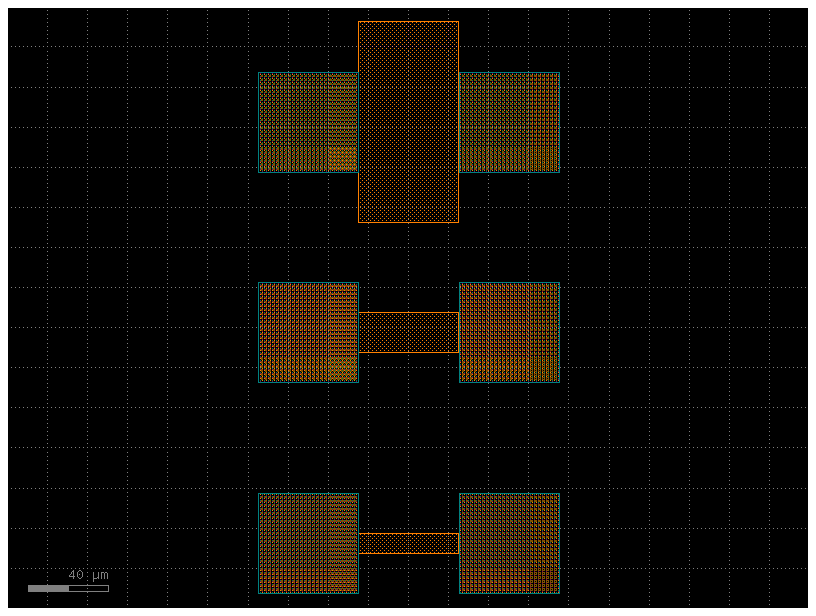
Load the layout¶
We import the GDS generated in notebook 1 to read the device geometry. Each instance in the top-level cell has cell.info.width and cell.info.length attributes set by resistance_sheet, so we can read the physical dimensions directly from the layout rather than hardcoding them.
@cache
def load_gds():
return gf.import_gds("resistance.gds", skip_new_cells=True)
gds = load_gds()
gds

Clean up existing files (optional)¶
Delete any files from a previous run of this notebook so you start fresh. The GDS uploaded in notebook 1 is preserved.
existing_files = client.query_files(tags=["project:tutorial_resistance", user])
to_delete = [f for f in existing_files if f["original_name"] != "resistance.gds"]
for file in tqdm(to_delete):
with suppress(RuntimeError):
client.delete_file(file["id"])
print(f"Deleted file {file["original_name"]}")
0%| | 0/138 [00:00<?, ?it/s]
Deleted file wafer_sheet_resistance.png
Deleted file wafer_sheet_resistance.png
Deleted file die_sheet_resistance.png
Deleted file die_sheet_resistance.png
Deleted file die_sheet_resistance.json
Deleted file die_sheet_resistance.json
Deleted file die_sheet_resistance.png
Deleted file die_sheet_resistance.json
Deleted file die_sheet_resistance.png
Deleted file die_sheet_resistance.json
Deleted file die_sheet_resistance.png
Deleted file die_sheet_resistance.png
Deleted file die_sheet_resistance.json
Deleted file die_sheet_resistance.json
Deleted file die_sheet_resistance.png
Deleted file die_sheet_resistance.json
Deleted file die_sheet_resistance.png
Deleted file die_sheet_resistance.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement_linear_fit.png
Deleted file iv_measurement_linear_fit.json
Deleted file iv_measurement.parquet
Deleted file iv_measurement.parquet
Deleted file iv_measurement.parquet
Deleted file iv_measurement.parquet
Deleted file iv_measurement.parquet
Deleted file iv_measurement.parquet
Deleted file iv_measurement.parquet
Deleted file iv_measurement.parquet
Deleted file iv_measurement.parquet
Deleted file iv_measurement.parquet
Deleted file iv_measurement.parquet
Deleted file iv_measurement.parquet
Deleted file iv_measurement.parquet
Deleted file iv_measurement.parquet
Deleted file iv_measurement.parquet
Deleted file iv_measurement.parquet
Deleted file iv_measurement.parquet
Deleted file iv_measurement.parquet
Deleted file iv_measurement.parquet
Deleted file iv_measurement.parquet
Deleted file iv_measurement.parquet
Deleted file iv_measurement.parquet
Deleted file iv_measurement.parquet
Deleted file iv_measurement.parquet
Generate and upload measurements¶
For each combination of wafer, die, and device instance we generate a synthetic IV curve and upload it. The cell tag uses the cell name from the GDS (which encodes the width), and width and length tags carry the physical dimensions in µm so downstream analysis can recover the sheet resistance formula without needing to re-read the layout. Because the dimensions are tags rather than parts of a filename, any downstream query can use query_files(tags=["width:10", "die:0,0"]) without parsing filenames or maintaining a lookup table.
sheet_resistance = 100 # Ω/sq
current_sweep = np.linspace(0, 10, 21) # mA
for wafer, die in tqdm(list(product(wafer_ids, dies))):
# Per-wafer multiplicative shift (±20%)
wafer_factor = 1 + 0.20 * (2 * np.random.rand() - 1)
die_id = f"{die['x']},{die['y']}"
for inst in gds.insts:
width_um = inst.cell.info.width
length_um = inst.cell.info.length
# Nominal resistance from sheet resistance formula: R = Rs × L / W
nominal_R = sheet_resistance * length_um / width_um
# Apply device-to-device variation (±5%)
device_factor = 1 + 0.05 * (2 * np.random.rand() - 1)
resistance = nominal_R * wafer_factor * device_factor
# Simulate defects: 5% open circuits, 3% short circuits
defect_roll = np.random.rand()
if defect_roll < 0.05:
resistance = 1e9 # open circuit
elif defect_roll < 0.08:
resistance = 0 # short circuit
voltage_mV = iv_resistance(resistance_ohm=resistance, current_mA=current_sweep)
# Add 5% multiplicative noise per sample
noise = 0.05 * (2 * np.random.rand(len(voltage_mV)) - 1) * voltage_mV
voltage_mV = voltage_mV + noise
data = pd.DataFrame(
{"current_mA": current_sweep, "voltage_mV": voltage_mV}
)
client.add_file(
data,
tags=[
user,
"project:tutorial_resistance",
f"wafer:{wafer}",
f"die:{die_id}",
f"cell:{inst.cell.name}",
f"device:{inst.name}",
f"length:{inst.cell.info.length}",
f"width:{inst.cell.info.width}",
],
filename="iv_measurement.parquet",
)
0%| | 0/8 [00:00<?, ?it/s]
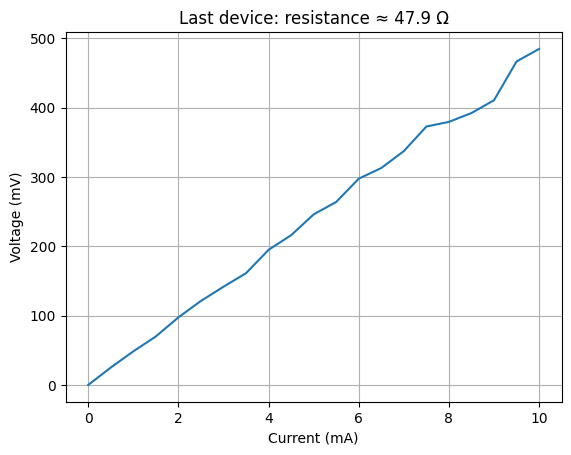
# Save one example measurement locally for use in notebook 3
data.to_parquet("last_measurement.parquet")
plt.plot(data["current_mA"], data["voltage_mV"])
plt.xlabel("Current (mA)")
plt.ylabel("Voltage (mV)")
plt.title(f"Last device: resistance ≈ {resistance:.1f} Ω")
plt.grid(True)
plt.show()
What's next?¶
You have uploaded IV curves for every device on every die across three wafers. The next notebook defines a linear_fit analysis function, wraps it in a pipeline that triggers automatically on new uploads, and runs it on all the files you just uploaded to extract resistance from each IV curve.